Our Solution
Fast drug discovery for pharma and biotechs
We partner with pharmaceutical companies and biotechs to provide a range of protein engineering and drug discovery services, based on the technology we develop.
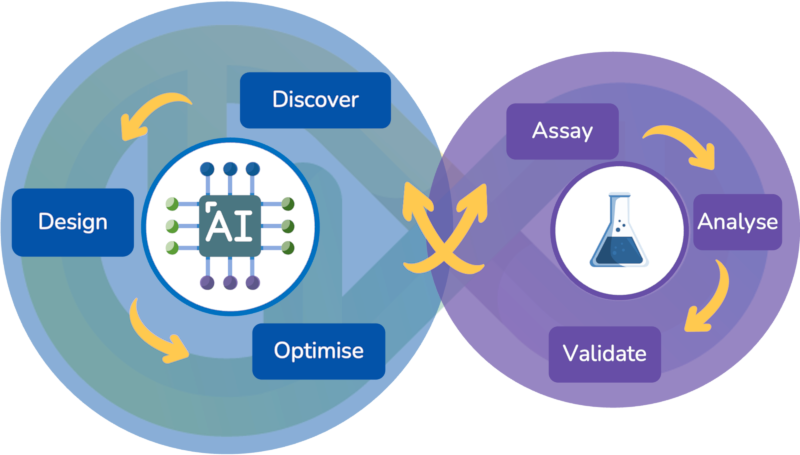
Our state-of-the-art AI/ML algorithms, coupled with in-silico pipelines and workflows, enables us to rapidly develop new therapeutic lead molecules.
Wet-lab based approaches to drug discovery can be slow and expensive, requiring many rounds of optimisation to yield results.
Our approach is different: we find high-quality candidates in silico, then work with you to validate them in the lab. Our technology allows us to find these candidates in days, rather than months or years as for traditional wet-lab approaches.