
The rapid advancement of AI technology and in-silico modelling of biologics are opening up new ways to design and discover biologics.
It is now entirely possible to design completely new de novo biologics on the computer or to have massive virtual screening campaigns to discover lead molecules much faster than what is possible in the lab.
At SilicoGenesis, the process of design, discovery and optimization of biologics starts in silico. Once lead molecules are identified, these molecules are then tested in the lab to verify their efficacy and properties.
Our process starts with the basics: we fully characterise the 3D structure of starting material for your drug discovery project including targets, hits and leads. If necessary, we identify hits via virtual screening or de novo design. We then perform hit-to-lead optimisation to produce candidates ready for testing and validation.
We characterise the structure of general proteins, antibodies, nanobodies and other biologics to identify protein-protein interaction sites, paratopes and epitopes. When necessary, we predict the 3D structure of proteins and antibodies from their sequence.
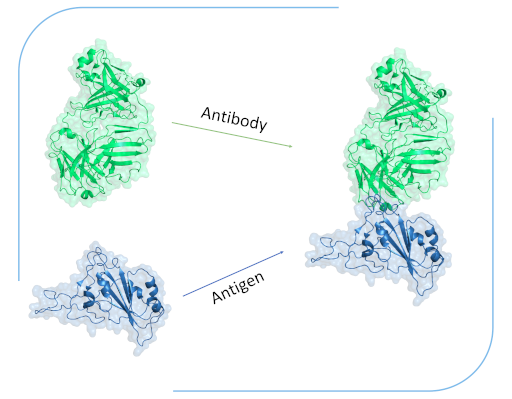
We find potential binders for your target through structure-based virtual screening experiments. Alternatively, we generate de novo candidates using generative machine learning models.
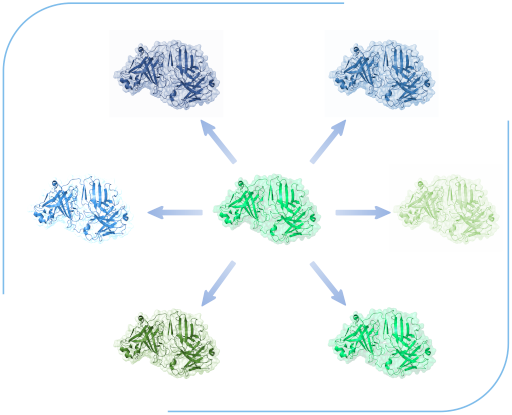
We optimise binders towards a desired affinity goal or species cross-reactivity threshold. These affinity-optimised binders can be validated for expression and functionality validations.
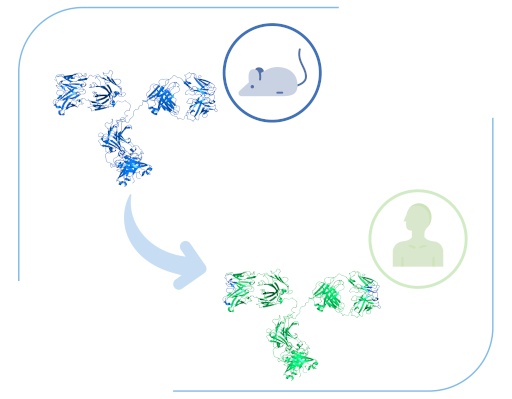
When antibodies have been sourced from non-human animal models, we perform humanisation and assess the “humanness” score of your antibody sequences without negatively altering their binding properties.
We prepare optimised binders for preclinical testing by critically assessing their structural properties and through mitigating sequence liabilities or developability issues.
We collaborated with a large European pharmaceutical company on an affinity maturation project for a difficult target. Given the sequence of the antibody candidate we predicted its structure, docked it to the target and performed in silico saturation mutagenesis.
Within 25 days we correctly identified all of the most affinity-enchancing mutations, confirmed by ELISA and SPR.
BioIncubator, Gaston Geenslaan 1, 3001 Leuven, Belgium
info@silicogenesis.com
+27 83 585 6050
Copyright © 2021-2023 SilicoGenesis. All Rights Reserved.
Leuven, Belgium and Johannesburg, South Africa.